Foundational questions on (cell) size determination in plants is the prime interest of the cluster group of Molecular Plant Physiology (MoPP). We particularly emphasize on differential control of plant growth from a subcellular to organ scale, deciphering plant architectural aspects that have importance for agriculturally important traits.
Building and conducting research projects is convoluted due to the complexity of biology and the ability of the actual people involved. While we clearly define the goals of our hypothesis driven projects, many decisions need to be taken on the go. Within the MoPP cluster group, we break up this complexity by sharing our team knowledge, jointly designing visions and solutions to deliver mechanistic knowledge on plant growth and adaptation.
Ongoing Projects:
Differential growth control along an organ defines its form and function. But how does it work?
To investigate differential organ growth, we focus on asymmetric growth regulation in lateral roots in response to gravity. Auxin controls the angular growth of lateral roots and thereby determines whether a root system deeply penetrates the soil or expands radially (Ruiz Rosquete et al., 2013; Ruiz Rosquete et al., 2018; Waidmann et al., 2019). We currently investigate how intrinsic and extrinsic cues shape this root architecture trait.
The size of the cell is defined by intracellular and extracellular processes. But how is this coordinated?
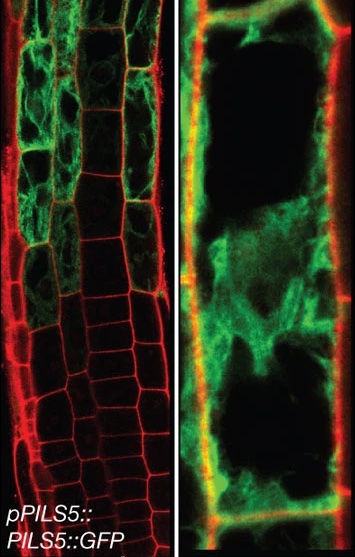
We investigate neighbouring root epidermal cell files that display distinct cell size control. We utilize these cell files, functionally facilitating the root-soil interface, as a cell-size determination model to monitor how internal and externalsignals get integrated (Löfke et al., 2013; Löfke et al., 2015; Scheuring et al., 2016; Dünser et al., 2019). We currently unravel environmental and cellular signals that define the size of the vacuole, which impacts on cell size determination by ensuring cytosol homeostasis during cellular expansion.
The phytohormone auxin is a central regulator of plant growth.
But how is auxin subcellularly controlled?
In an in-silico screen, we have identified the ER resident PIN-LIKES (PILS) family of auxin transport facilitators (Barbez et al., 2012). PILS proteins control intracellular auxin accumulation at the ER and restrict nuclear availability and signalling of auxin, presumably by limiting auxin diffusion into the nucleus (Barbez et al., 2012; Feraru et al., 2012; Beziat et al., 2017; Feraru et al., 2019; Sun et al., 2020). PILS proteins and PILS gene regulation is highly sensitive to environmental perturbations, which allows the integration of external cues into growth programs, shaping plant adaptational responses (Beziat et al., 2017; Feraru et al., 2019; Sun et al., 2020).