Foundational questions on (cell) size determination in plants is the prime interest of the cluster group of Molecular Plant Physiology (MoPP). We particularly emphasize on differential control of plant growth from a subcellular to organ scale, deciphering plant architectural aspects that have importance for agriculturally important traits.
Building and conducting research projects is convoluted due to the complexity of biology and the ability of the actual people involved. While we clearly define the goals of our hypothesis driven projects, many decisions need to be taken on the go. Within the MoPP cluster group, we break up this complexity by sharing our team knowledge, jointly designing visions and solutions to deliver mechanistic knowledge on plant growth and adaptation.
Ongoing Projects:
Differential growth control along an organ defines its form and function. But how does it work?
To investigate differential organ growth, we focus on asymmetric growth regulation in lateral roots in response to gravity. Auxin controls the angular growth of lateral roots and thereby determines whether a root system deeply penetrates the soil or expands radially (Ruiz Rosquete et al., 2013; Ruiz Rosquete et al., 2018; Waidmann et al., 2019). We currently investigate how intrinsic and extrinsic cues shape this root architecture trait.
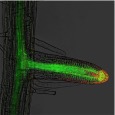
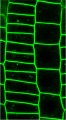
The size of the cell is defined by intracellular and extracellular processes. But how is this coordinated?
We investigate neighbouring root epidermal cell files that display distinct cell size control. We utilize these cell files, functionally facilitating the root-soil interface, as a cell-size determination model to monitor how internal and externalsignals get integrated (Löfke et al., 2013; Löfke et al., 2015; Scheuring et al., 2016; Dünser et al., 2019). We currently unravel environmental and cellular signals that define the size of the vacuole, which impacts on cell size determination by ensuring cytosol homeostasis during cellular expansion.
The phytohormone auxin is a central regulator of plant growth.
But how is auxin subcellularly controlled?
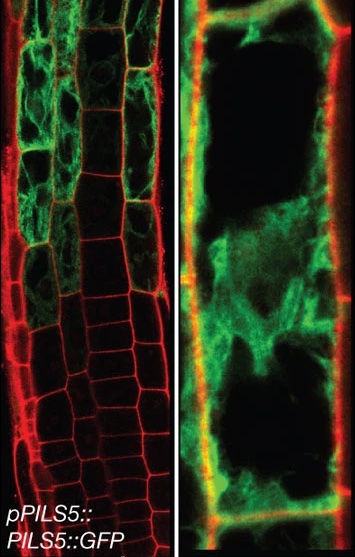
In an in-silico screen, we have identified the ER resident PIN-LIKES (PILS) family of auxin transport facilitators (Barbez et al., 2012). PILS proteins control intracellular auxin accumulation at the ER and restrict nuclear availability and signalling of auxin, presumably by limiting auxin diffusion into the nucleus (Barbez et al., 2012; Feraru et al., 2012; Beziat et al., 2017; Feraru et al., 2019; Sun et al., 2020). PILS proteins and PILS gene regulation is highly sensitive to environmental perturbations, which allows the integration of external cues into growth programs, shaping plant adaptational responses (Beziat et al., 2017; Feraru et al., 2019; Sun et al., 2020).
Phloem differentiation and function – Lothar Kalmbach
Multicellular organisms like plants, animals, and humans rely on vascular systems to distribute water, nutrients, and signaling molecules between distant body parts. In the plant vascular system, the xylem transports water and mineral nutrients from the root to the aerial organs and the phloem to transport water and organic compounds like sugars from photosynthetically active sources (i.e., mature leaves) to non-photosynthetic or heterotrophic sinks (i.e., roots, fruits, and buds). This long-distance transport needs to be reliable, efficient, and adaptable.
Key to the physiological function of the phloem are its conductive cells, the sieve elements that together make up the sieve tube, and its immediate neighbors, companion cells and phloem parenchyma cells (in leaves) or phloem pole pericycle cells (in roots), that either load organic compounds into the sieve tube in source tissues or unload the phloem sap in sinks. Sieve elements are highly specialized for high-volume transport: During differentiation, they degrade their nucleus and most organelles (Furuta et al., 2014), undergo considerable cell wall thickening (Truernit et al., 2008) and develop large sieve pores to connect neighboring cells (Kalmbach and Helariutta, 2019). While many features of these cells have been anatomically described already decades ago, the molecular and genetic factors that control this dramatic cellular differentiation are largely unknown.
Ongoing Projects:
- Regulation of tissue-specific callose deposition
- Callose deposition is an important intermediate step in sieve element differentiation that enables formation of bigger sieve pores allowing higher phloem transport. We recently localized CALLOSE SYNTHASE 7 as the first sieve pore localizing protein providing a tool to both visualize and genetically dissect sieve pore formation in the model plant Arabidopsis thaliana (Kalmbach et al., 2023). Combining cell biology, candidate gene approaches, Crispr-Cas9 gene editing, and physiological assays, we are identifying general as well as tissue-specific mechanisms that control callose deposition. This has broad impact beyond phloem differentiation as callose is rapidly incorporated in the cell wall during cell division, upon physical damage, or pathogen attack and therefore an important cell wall modification for plant growth and defense.
- Tissue-specific pectin signaling in growth and development
- Oligogalacturonides (OGs) are small breakdown products that are released by secreted pectin-degrading enzymes from pathogens and elicit strong defense responses in plants. Plants themselves produce a variety of pectin modifying enzymes, which also produce OGs, some of which are very specifically expressed and critical to developmental processes. Recently, we identified the pectate lyase PLL12 that modifies pectin specifically in the phloem in a tightly defined developmental window. While necessary in the developing phloem, strong ubiquitous expression provokes all hallmarks of a strong pathogen response and is lethal (Kalmbach et al., 2023). Additionally to regulating phloem sieve pore diameter, PLL12 therefore likely has additional roles to produce OGs in a tissue-specific context to regulate phloem developmental processes. We are currently developing PLL12 and phloem sieve element differentiation as a model to study how plants use cell wall breakdown products beyond and independent of pathogen-induced defense reactions. To this end, we establish a suite of tools to map and manipulate production of and responses to OGs at cellular resolution. This will identify mechanisms how plants use cell wall breakdown products for defined developmental decision independent of and beyond pathogen-induced defense reactions.
References
- Furuta, K.M. et al. (2014). Arabidopsis NAC45/86 direct sieve element morphogenesis culminating in enucleation. Science 345: 933–937.
- Kalmbach, L. et al. (2023). Putative pectate lyase PLL12 and callose deposition through polar CALS7 are necessary for long-distance phloem transport in Arabidopsis. Curr Biol 33: 926-939.e9.
- Kalmbach, L. & Helariutta, Y. (2019). Sieve Plate Pores in the Phloem and the Unknowns of Their Formation. Plants (Basel) 8.
- Truernit, E. et al. (2008). High-resolution whole-mount imaging of three-dimensional tissue organization and gene expression enables the study of Phloem development and structure in Arabidopsis. Plant Cell 20: 1494–1503.